
A new method to study specific changes in DNA after replication has been published as a technical report in Nature Cell Biology. Researchers developed a highly sensitive, quantitative mass spectrometry-based approach, called iDEMS (isolation of DNA by EdU labelling for Mass Spectrometry).
“The novelty in our work is that we didn’t use sequencing methods widely used in this field, instead we used mass spectrometry, which is the first time this approach has been used to measure DNA modifications on purified, replicated DNA,” says Dr Stewart-Morgan, co-first author of the report, from the Groth laboratory at the Novo Nordisk Foundation Center for Protein Research (CPR) at University of Copenhagen.
This unique approach is the result of a joint project with the Hajkova laboratory at MRC London Institute of Medical Sciences (LMS). “In the Groth laboratory we have expertise in replication and the Hajkova laboratory has expertise in studying DNA methylation by mass spectrometry. I think this multidisciplinary collaboration is a large part of the reason why the project has been so successful,” Dr Stewart-Morgan explains. “The results of our research using iDEMS are definitive and open new avenues for future research.”
DNA modifications and cell stability
The genome is the entire set of DNA instructions found in a cell. Virtually all cells in an organism contain the same genetic information – but which genes are expressed is based on the cell’s function. This cell-specific gene expression is regulated by the cell’s epigenome, which consists of proteins bound to DNA, as well as direct chemical modifications to DNA. One of the most important epigenetic regulators is DNA methylation – a chemical marker which turns off regions of the genome that should not be expressed. The pattern of these markers is very important in maintaining a cell’s stability and identity: for example, DNA methylation in a liver cell will differ from the DNA methylation pattern in a blood cell.
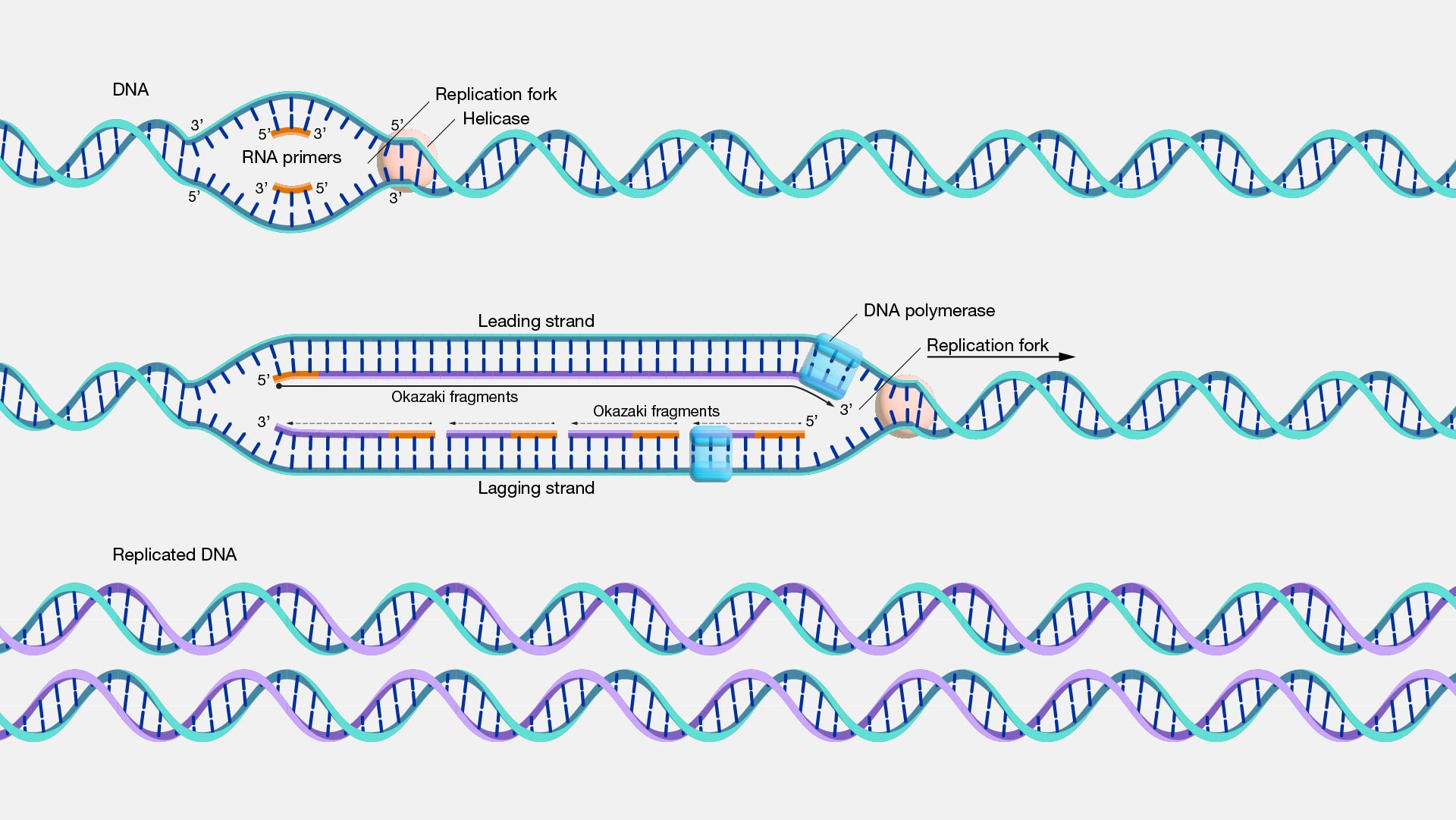
When DNA is replicated during cell division, the epigenetic marks associated with the DNA, including DNA methylation, are diluted. The newly created DNA strands need to re-establish the level and pattern of methylation to maintain control of gene expression, genomic stability and the epigenetic memory of the cell’s identity.
However, much about this process is unknown, and loss of DNA methylation is a common feature in cells that have divided many times, such as cancer cells which are very proliferative and aged cells that have replicated many times over the course of a person’s lifespan. In recent years several groups have tried to investigate this process using sequencing methods, however the exact kinetics of post-replicative methylation maintenance remained unclear.
Methylation re-establishment
Using iDEMS, the researchers found that DNA methylation levels increase steadily after replication, and after 4 hours the levels on replicated DNA and the genomic DNA were equal. This indicates that this process proceeds at a steady, slow pace. However, it is outpaced by cell division.
“Over time cells don’t have long enough to re-establish their methylation after replication, and the methylation of the genome is eventually diluted. This is the first time very clear kinetics for methylation re-establishment have been shown. Furthermore, we saw absolute quantification of the levels of DNA methylation, enabling us to distinguish which methylation marks were newly established. This gave us confidence in our kinetic measurements,” Dr Stewart-Morgan reports.
A second chemical marker
The researchers also used iDEMS to study a second marker – DNA hydroxymethylation – which is a much rarer genomic marker than methylation. Their results corroborated earlier research, says Dr Stewart-Morgan: “We found that one DNA strand, the template or ‘parental’ strand, always has more hydroxymethylation than the other ‘daughter’ strand, supporting earlier work which indicated that this marker distinguishes DNA strands based on age,” she says.
“However, we also discovered that there is no point at which the levels of hydroxymethylation are equal between the parental and daughter strands throughout the cell cycle. This opens new questions about how this difference between strands may be used by cells, for example during DNA repair.”
The potential of iDEMS
By directly quantifying DNA modifications on replicated DNA, iDEMS resolves DNA methylation and hydroxymethylation kinetics following DNA replication. “iDEMS is a dynamic and informative tool for addressing important questions in epigenome maintenance and DNA modification biology,” Dr Stewart-Morgan says.
Looking to the future, iDEMS will be useful in profiling methylation and hydroxymethylation dynamics in different cellular contexts, including ageing and cancer evolution. Compared with sequencing data, mass spectrometry provides a simple, fast readout, and iDEMS could therefore be useful where efficiency is key, such as in medical settings and drug discovery studies.
“Our results highlight how important new methods are for understanding biology through more than one lens. iDEMS is extremely flexible, as it can be combined with other established methods used in molecular biology to look at the epigenome. This method therefore adds an important tool to the suite of technologies investigating epigenome stability,” concludes Dr Stewart-Morgan.

