
Researchers at Cardiff University have used x-ray crystallography and computer simulation to get a closer look at how viruses bind cells and cause infection.
The new insight could help in the development of drugs and therapies for infections and further advance the exploitation of viruses for medical treatments.
The first author of the study, Alex Baker from Cardiff University’s School of Medicine, said, “We were interested in developing viruses for therapeutic applications, such as treating cancers, and for vaccine applications.
We focussed on two specific viruses, called Ad26 and Ad48. These viruses are in clinical trials where they are showing promise as vaccines to protect against Ebola virus and HIV infection. We wanted to know more about how these viruses work as vaccines.”
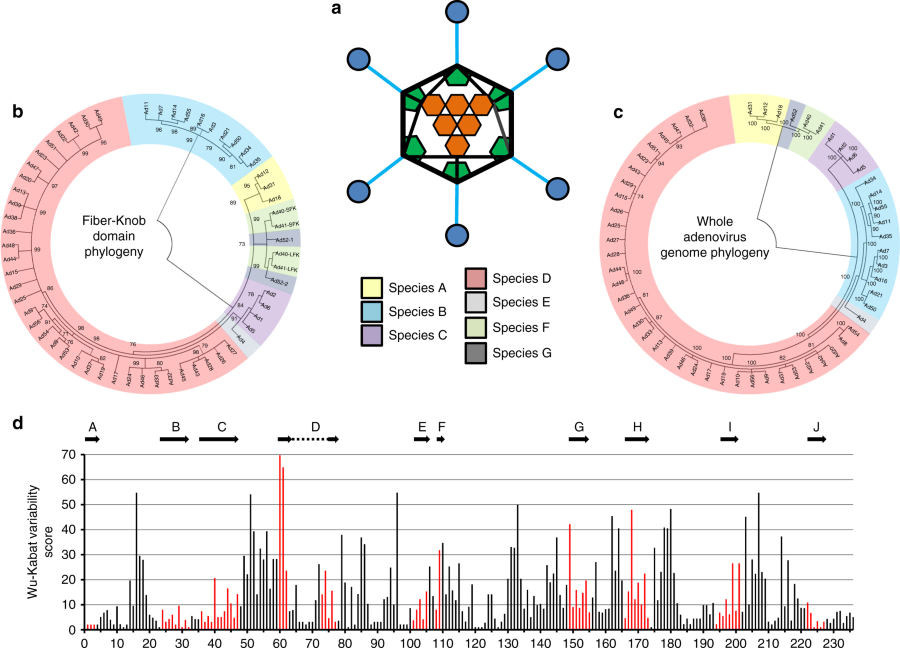
The team made very pure preparations of the virus protein that binds the virus to a cell during infection. Using this purified protein, they produced crystals and performed X-ray diffraction studies, which gave them an extremely accurate picture of the protein that the virus uses to bind and infect cells.
“We were able to investigate exactly how the viruses attach to proteins on the cell surface and we were surprised to find that they could not bind to a protein called CD46, which had previously been reported as the main virus receptor,” added Baker.
“Instead, we show that these viruses can bind weakly to a different entry receptor, called CAR. In so doing we identified a previously undiscovered mechanism that adenoviruses use to adjust their attachment to CAR.”
The senior author, Dr. Alan Parker from Cardiff University’s School of Medicine, said, “To train viruses into useful medicines, such as agents for treating cancer, the first step is to understand at the molecular level how these viruses work. This allows us to unpick the virus’s natural biology, and ‘tailor’ them into therapeutically useful agents.
“This study is important since it provides molecular level detail about the basic biology of the viruses we are manipulating. This information will help determine what the best antivirals might be to treat outbreaks of these viruses in their natural, disease causing state, but will also help progress their development for therapeutic purposes in the future.”
The research “Diversity within the adenovirus fiber knob hypervariable loops influences primary receptor interactionsis” is published in Nature Communications, and was funded by Tenovus Cancer Care, Cancer Research UK and Cancer Research Wales.
The study also involved collaborators based in Oxford and the Icahn School of Medicine at Mount Sinai in New York.

